About
Research Interests:
1. Cancer Metabolism
(i) We currently researching the metabolic interplay between exogenous cystine and glutamine dependence in triple-negative breast cancer, with view to finding new anti-cancer targets. https://www.nature.com/articles/s41420-025-02714-3
(ii) One of our recent Marie Curie funded research projects demonstrated a role for Interleukin-6 (IL-6) in anoikis resistance (an index of metastases) in oral squamous cell carcinoma (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9050791/pdf/12032_2022_Article_1664.pdf) and bioenergetic reprogramming on these cancer cells (https://doi.org/10.26124/bec:2022-0011). The work indicates that anti-IL6 receptor antibodies may be efficacious in stopping the spread of oral cancer.
(iii) We have also identified Nicotinamide N-Methyltransferase (NNMT) as a potential target in oral squamous cell carcinoma. https://iadr.abstractarchives.com/abstract/irish-iadr23-3978233/nicotinamide-n-methyltransferase-nnmt-is-an-anti-cancer-target-in-cultured-human-oral-squamous-cell-carcinoma-oscc
2. Cancer Cachexia
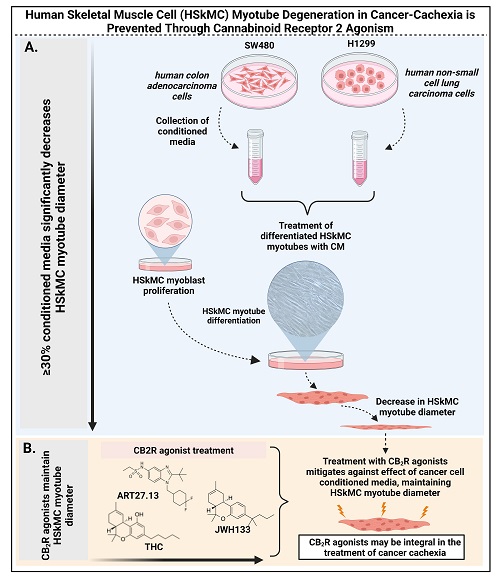
Our laboratory have established an in vitro primary human skeletal muscle cancer cachexia model. This model is being used to (i) identify anti-cachexia targets, (ii) test anti-cancer cachexia drug efficacy and (iii) identify protagonists causing cachexia. See our recent publication: “Cancer-Cachexia-Induced Human Skeletal Muscle Myotube Degeneration is Prevented via Cannabinoid Receptor 2 Agonism in Vitro,” (https://www.mdpi.com/1424-8247/16/11/1580/pdf) is published in the journal Pharmaceuticals. [November 2023].