Genomic Analysis in Yeast Reveals New Insights into Cellular Quiescence
13th February, 2017
It has been estimated that many cells spend the majority of their lifetime in a non-dividing, or quiescent state. During quiescence (or G0), these cells are often considered as ‘resting’, whereby metabolic activity and gene expression are reduced. Another key characteristic of quiescence is that this state is reversible, and cells can re-enter the cell cycle to resume growth when specific conditions occur.
Numerous cells in the human body persist in this non-dividing state, including neurons and muscle cells. Stem cells also form quiescent populations which, for example, can re-enter the cell cycle in response to tissue damage to aid repair. Furthermore it is now known that cancer cells can become quiescent, during which time they are more resistant to drug treatments. However, research into this so-called ‘inactive’ cellular state has often been over-looked, and the mechanisms that regulate quiescence are poorly understood.
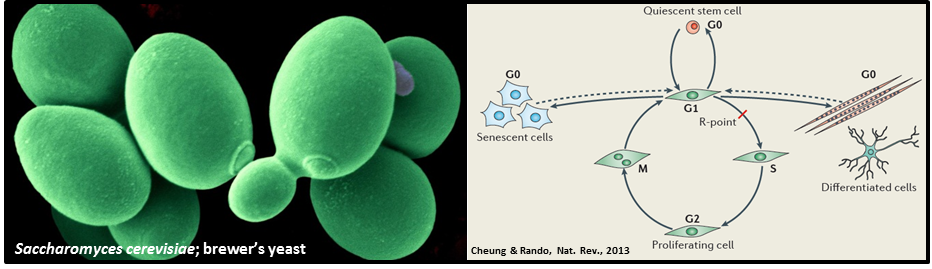
A collaboration between Dr Alastair Fleming (Department of Microbiology), Dr Karsten Hokamp (Department of Genetics) and Prof Mary Ann Osley at the University of New Mexico, USA, has used the humble brewer’s yeast, Saccharomyces cerevisiae, as a model organism in which to study cellular quiescence. The study mapped and compared the gene transcription machinery, and distinct chromosomal modifications that are associated with gene activity, across the entire genomes of actively-growing and quiescent yeast cells. Surprisingly, the study revealed that the genomes of the ‘inactive’ quiescent cells retained many of the chromosomal signatures that are normally associated with actively growing cells, despite their shut-down in cellular activity. Furthermore the transcription machinery, although inactive, remained associated with most genes across the quiescent cell genome. Together, the study revealed that quiescent yeast cells are not inert, but are highly poised to resume growth and proliferation when the environment becomes favourable.
Dr Conor Young, a lead author of the study said, ‘the most interesting aspect of the study was that yeast showed a burst of transcription activity just before their exiting the cell cycle, during which many of the chromosomal marks required for active growth were deposited along the cell genome. It is like the yeast cells make a ‘last-gasp’ attempt to arm themselves with the chromosomal attributes required to best ensure their survival and re-entry into the cell cycle’.
Assistant Professor of Microbiology, and co-senior author of the study, Dr Alastair Fleming stated, ‘This was an important study which mapped the chromosomal landscape of quiescent cells, and showed how this landscape was established. Our future aim is to uncover how the chromosomal modifications we have described contribute to the process of quiescence. By understanding the correct pathway cells take into and out of quiescence, it will be possible to identify where cells have deviated from this pathway during disease, which may yield future therapeutic targets’.
The study, funded by Science Foundation Ireland and the NIH, was published in the open-access BMC Genomics journal, and can be accessed here (10.1186/s12864-017-3509-9).
Young, Conor P. et al. ‘Distinct histone methylation and transcription profiles are established during the development of cellular quiescence in yeast’. BMC Genomics. 2017;18:107. doi:10.1186/s12864-017-3509-9.